Electrons have two intrinsic properties, charge and spin. The first is widely known, the second, less appreciated. The spin of an electron is a purely quantum mechanical property; it specifies the electron’s intrinsic angular momentum. A common classical analogy is to describe it in terms of a spinning top. Like a top, which can rotate clockwise or counterclockwise, an electron has two spin states, spin up and spin down. These two states have half-integer values of ℏ/2 and –ℏ/2. They have the symmetry properties of fermions and obey the Pauli Exclusion Principle: two electrons cannot be in the same quantum state. While classical electrostatics does not allow two electrons to occupy the same region of space in a stable system, it is possible in quantum mechanics as long as their spins are different. This principle guides the organization of electrons in atoms, as well as in the chemical bonds between them.
In biology, the charge of the electron is important due to its role in defining the Coulomb forces that govern interaction energies between molecules. Electrons also play essential roles in the transduction of energy and information. For example, in photosynthesis, light energy—the absorption of one or more photons—initiates electron transfer, which ultimately results in the production of adenosine triphosphate (ATP) molecules, a key source of energy for biological processes. The conventional picture considers only the electrons’ charge; the spin property is ignored and assumed to be unimportant. Recent work, however, indicates otherwise.1
Here we discuss recent discoveries pointing to the importance of spin in biological systems, such as its role in long-range electron transfer, in reactions that involve multiple electrons, and in biorecognition. All these phenomena arise from chiral-induced spin selectivity (CISS), which links electron transfer through chiral molecules to a specific spin state of the electron. When electrons are transferred through chiral molecules, one spin state of the electron is preferred.
Chirality is the name for the particular kind of symmetry that arises when one object is the mirror image of the other. An example is the human hand. Mirror images require no internal symmetries. Our two hands are called enantiomers of one another. Chiral molecules are referred to as being a right-handed enantiomer or a left-handed enantiomer.
Chiral Biomolecules
The basic molecular building blocks of life—DNA and proteins, for example—lack mirror symmetry. All organisms, plants, animals, and of course human beings, are made from molecules with “helical” symmetry in the shape of a screw.2 The components from which proteins are made—the amino acids—are enantiomerically pure in all organisms, and this purity is preserved with extremely high fidelity. Such enantiomeric selectivity is very difficult to achieve artificially in the laboratory. Moreover, the enantiopurity of drug medications is known to be essential to their efficacy. And without it, side effects are common and sometimes deadly.
For a long time, scientists have discussed the role of chirality in nature. Of particular interest have been questions such as, “Why do biological systems use only one enantiomer of a given biomolecule?” and, “Why is a particular enantiomer preserved so persistently through evolution?”3 One argument given for the existence of chiral molecular building blocks is that, in general, they are complex enough to enable the biochemical complexity of life, and, in particular, permit the construction of the necessary three-dimensional architectures.4 While this may be true, it does not explain the high fidelity of enantiorecognition in nature.
Long-Range Electron Transfer
A typical bond length in a molecule is 1 to 2 angstroms, yet natural biological processes move electrons from ten to a hundred times further in a single reactive step. While fully saturated organic molecules confine electrons to the length scale of a chemical bond, electrons in conjugated molecules can delocalize over many atomic centers. But biological systems do not use such molecules to transfer charge over long distances. Instead biological electron transfer processes use proteins, which are chiral and are comprised largely of saturated bonds. Why?
The CISS effect provides a possible answer. When electrons move elastically through chiral molecules, their spin and linear momentum are coupled; they cannot be reflected back without also flipping their spin, an improbable event in organic molecules. An electron should be able to propagate farther through a chiral molecule than through an achiral analog. This feature enhances the efficiency of electron transfer through chiral proteins.
It is important to appreciate that molecules in a biological environment undergo significant thermal fluctuations. Nuclear and electronic motions vary and electron transfer thus occurs under the influence of an electronic potential that fluctuates. Under such conditions, backscattering could be significant if the CISS effect did not constrain the process. The use of chiral molecules as bridges for electron transfer could provide symmetry constraints that enhance electron transfer efficiency. Indeed, recent experiments reveal that spin plays a role in electron transfer through peptides, proteins, and DNA.5 For example, the electrons transferred through Photosystem I are spin-polarized.6
Multi-Electron Processes
Many biochemical processes, such as oxygen formation in photosynthesis and respiration, require the transfer of multiple electrons. In the photosynthetic system, water can be split to generate oxygen and protons with a quantum efficiency of about 90%.7 This is much higher than artificial clean energy sources.8 Water electrolyzers and photodriven electrolysis have been demonstrated and such devices are commercially produced, but to implement them broadly will require additional significant improvements in efficiency.
There are two major limitations on the artificial photodriven formation of hydrogen. Because of slow kinetics for oxygen evolution at the photoanode, a high overpotential (circa 0.5V) is required to initiate the reaction. Second, the non-selectivity of the oxidation results in the production of hydrogen peroxide. In addition to improving device efficiencies, overcoming these limitations in artificial systems may provide insights into analogous bioprocesses.
In splitting water, two hydroxide ions are oxidized on an anode to form two OH radicals. These, in turn, react to form an oxygen molecule in its triplet electronic ground state and two protons. The protons are then reduced on a cathode, and hydrogen molecules are formed. Recent work shows that a chiral anode provides more efficient water splitting than an achiral one, presumably because it aligns the unpaired spins of the OH radicals, enhancing the yield of triplet oxygen molecules.9 This interpretation explains why adsorbing chiral molecules on the anode both lowers the overpotential and reduces hydrogen peroxide production.
Enantioselective Biorecognition
Biorecognition events are often modeled using a force field scheme—based on the foundational work of the late Shneor Lipson—for describing the interaction between biomolecules. Current methods use only the charge of the electron; its spin is ignored.10 This classical picture is adequate as long as the electrons are not in close proximity. When molecules share electron clouds, spin and its symmetry constraints become important. Current methods do not properly account for the enantioselectivity and binding energies in biorecognition unless they are corrected empirically. These models may be missing an essential feature.11
A recent study has suggested a way to bridge this gap.12 An electric field was applied across a monolayer of adsorbed molecules to redistribute the charge in the film and then, using the Hall effect, the magnetization of the film was measured. This magnetization depended on the molecule’s handedness and its length, and resulted from the spin polarization of the electron cloud. Charge polarization in chiral molecules, the experiment indicated, is accompanied by spin polarization. The spin polarization imposes a symmetry constraint from the Pauli exclusion principle that affects the electron cloud overlap. As two chiral molecules interact, they induce a charge redistribution and a spin polarization in their electronic clouds which change the interaction energy. Current force field models fail to account for such effects.
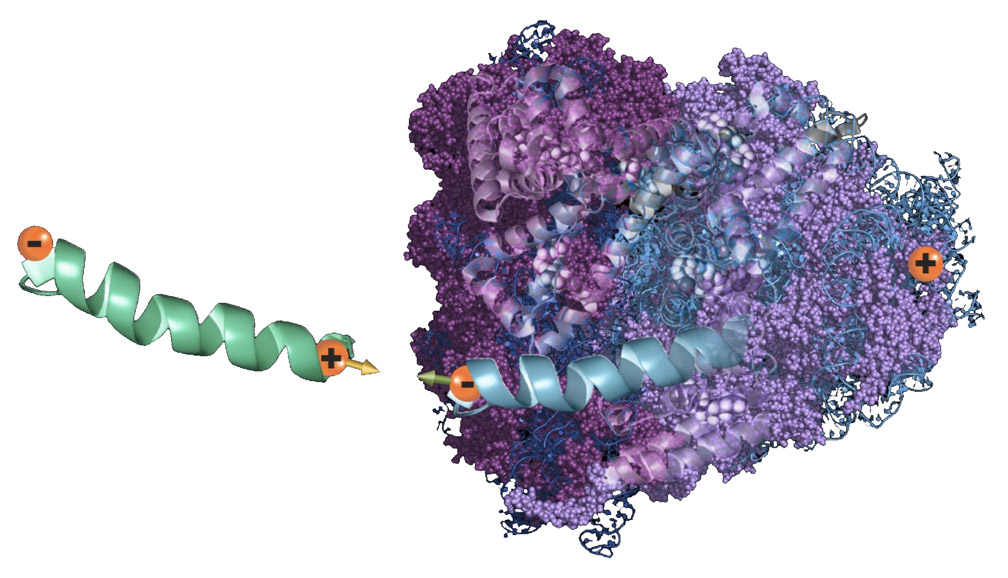
Consider the following thought experiment. A relatively small chiral molecule interacts with a system that contains chiral objects (Figure 1). The electron clouds on the two interacting bodies are polarized and generate a dipole moment at each of the interacting species. As the charge polarization evolves, a spin polarization (orange arrows) arises, which affects the interaction energy, depending on which enantiomers are involved. For two enantiomers with the same handedness, their spins point in opposite directions at the interaction region. The exchange interaction between the molecules—resulting from the overlapping of their wave functions—has a singlet (spin-paired) arrangement and satisfies the Pauli exclusion principle. If, on the other hand, the two enantiomers have opposite chirality, their spins at the interaction region will point in the same direction and the spin polarization of the overlapping electron clouds will be more like a triplet (spin aligned). The wave function overlap will be more repulsive. The interaction energy between the molecules in the overlap region differs intrinsically according to the chirality. The interaction is inherently enantioselective.
Figure 1.
The interaction between a small chiral molecule (green) and a chiral system (purple). When the two species interact via dispersive forces, the interaction induces an asymmetry in the electrons’ distribution resulting in an induced dipole in each of the interacting systems. This charge polarization is accompanied by a spin polarization. When the two species are of the same handedness, the spin polarization (orange arrows) is such that the electrons in the overlapping electron clouds have spins opposite to each other; the interaction is like a singlet state. When the two interacting molecules are of opposite chirality, the overlapping electron clouds have spins parallel to each other, like a triplet.
This thought experiment was considered more quantitatively by calculating the interaction between two methyl groups covalently linked to an asymmetric carbon, namely R—CH3…CH3—R. Here R is a chiral –CFClBr group that can be either R or S handed. The molecules and geometry of the interaction were chosen so that any difference in the interaction energy would arise solely from the symmetry constraints introduced by the spin polarization, which accompanies the charge polarization. The calculations showed that the interaction is less repulsive—by about 0.5 kcal/mol at 2.6 angstroms—for anti-parallel spins than for parallel ones. Distances associated with substrate-protein interactions are often under 3 angstroms; this phenomenon can thus be significant.13 Moreover, the effect need not be directly at the site of the asymmetric carbon, it can be remote; multiple contact points can contribute to enantiospecificity.
While it is well known that electron exchange and charge penetration contribute significantly to intermolecular forces at short range, they are still described by a simple r–12 repulsive potential, using a Lennard-Jones model.14 Consideration of the electron spin, however, suggests that a new term needs to be included in force field models, or that existing repulsive terms should be modified, in order to account for the different exchange energies experienced by molecules of different handedness.
Spin polarization, which in chiral molecules accompanies charge polarization, is another way quantum mechanics affects biology. The coupling of the spin of an electron to its motion in chiral molecules and the resulting CISS have significant effects, even at physiological temperatures. For biomolecules, the effects are wide ranging, from the efficient transfer of electrons over relatively long distances, to enhanced selectivity of oxidation reactions, to the efficiency of enantioselective biorecognition.
Chiral symmetry and CISS bear some resemblance to topological effects in physics.15 It is interesting to note that the 2016 Nobel Prize in physics was given to Duncan Haldane, Michael Kosterlitz, and David Thouless, for “theoretical discoveries of topological phase transitions and topological phases of matter.”16
Much remains to be discovered.17